Genomic observatories
Under the genomic observatories, we investigate a range of habitats and taxa in the Belgian Part of the North Sea using DNA-based techniques. In comparison to traditional morphological analysis, DNA-based techniques allow faster screening of samples without the need for in-depth taxonomic expertise, and in some cases have proven to be more cost-efficient as well. Genomic observatories offer opportunities for the early detection of non-indigenous species, filling gaps in current regional observations and have a high level of standardisation across regions.
Genomic observatories
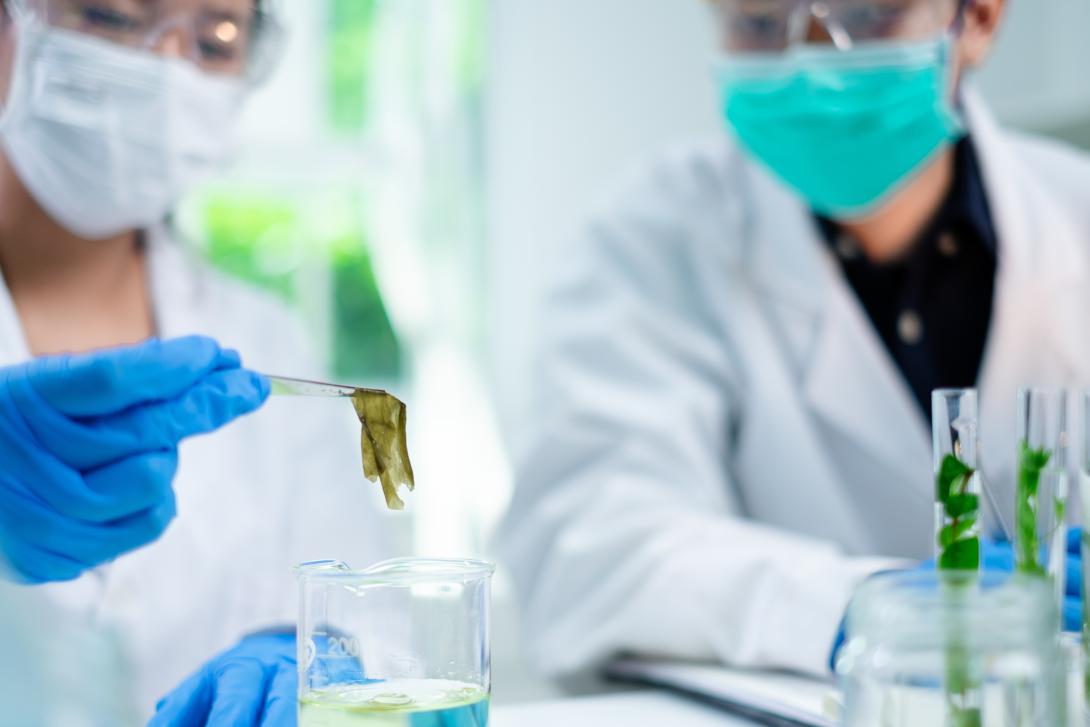

Traditional morphological monitoring is time-consuming, expensive and requires taxonomic experts who can be hard to find. The use of DNA-based techniques allows for faster screening of samples, without the need for in-depth taxonomic knowledge and can be standardised across regions. In many cases, the method has proven to be more time- and cost-efficient than traditional analysis, allowing for a larger spatial and temporal coverage of monitoring.
Typically, two types of DNA-based methods are used in monitoring. Current monitoring largely uses metabarcoding of bulk samples, samples containing the groups of interest. A more novel technique makes use of eDNA, or DNA that can be extracted from environmental samples like water or sediment samples.
This includes two types of DNA; 1. DNA from microscopic taxa present in the water or sediment samples themselves and 2. extra-organismal DNA originating from e.g. shed skin/tissues, excretion products, cell lysis or biologically active propagules. The latter allows us to detect the presence of a species without a specimen of the species being in the sample itself, in essence it is detected through the environment.
DNA-based methods allow us to cover a range of different taxa, including rare, cryptic or elusive taxa that are otherwise sometimes missed by traditional sampling and analysis. eDNA analysis also allows for non-invasive sampling of sensitive and rare and/or endangered taxa without putting pressure on the population.

Infrastructure
In the Belgian Part of the North Sea (BPNS), DNA and eDNA is sampled during the monthly LifeWatch campaigns by taking water samples. The grid of stations that are visited on a monthly and seasonal basis allow for a good spatio-temporal coverage of the BPNS.
LifeWatch also facilitates various other genomic monitoring events in collaboration with international consortia and Research Infrastructures like Ocean Sampling Day (OSD), EMO BON (EMBRC), ARMS-MBON (EMBRC, AssemblePlus) and Jerico.
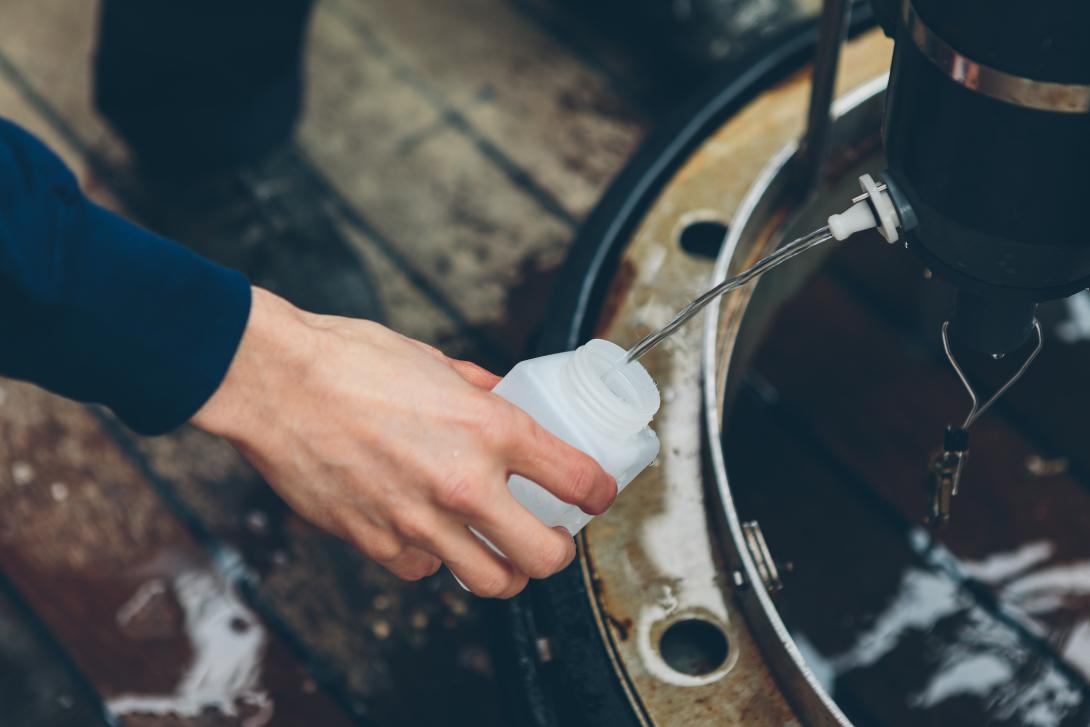
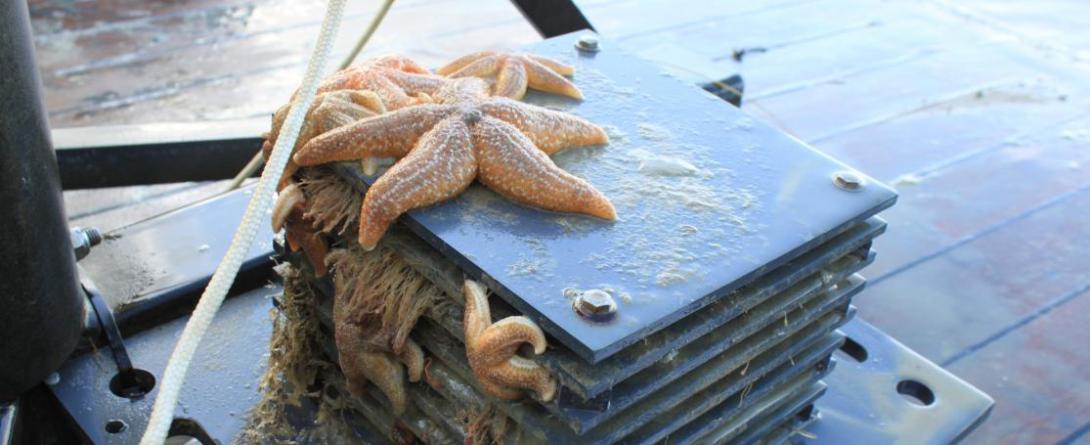


Collected water samples are filtered onboard and stored for further processing in line with the Ocean Sampling Day protocol for analysis of microbial and plankton communities. Sequencing is performed at a centralised facility, and lab and sampling protocols are standardised.
Under EMO BON, dedicated water column and soft sediment samples are collected bi-monthly at station 330, which was selected for its long-standing biodiversity observations. EMO BON uses heavily standardised protocols for sampling and analysis and provides centralised sequencing on the European scale. Through EMO BON we target plankton communities through water column samples and microbial, meiobenthos and macrobenthos soft sediment communities in the BPNS. Through the EMO BON network, Belgium is one of the 16 connected observatories over Europe and the Mediterranean.
ARMS-MBON enables monitoring of hard substrate fauna by deploying Autonomous Reef Monitoring Reefs (ARMS) units at various locations at the BPNS during one year long deployments. The ARMS MBON project has 20 observatories all over Europe and the Arctic and provides metabarcoding of samples to its partners targeting COI and 18S markers.
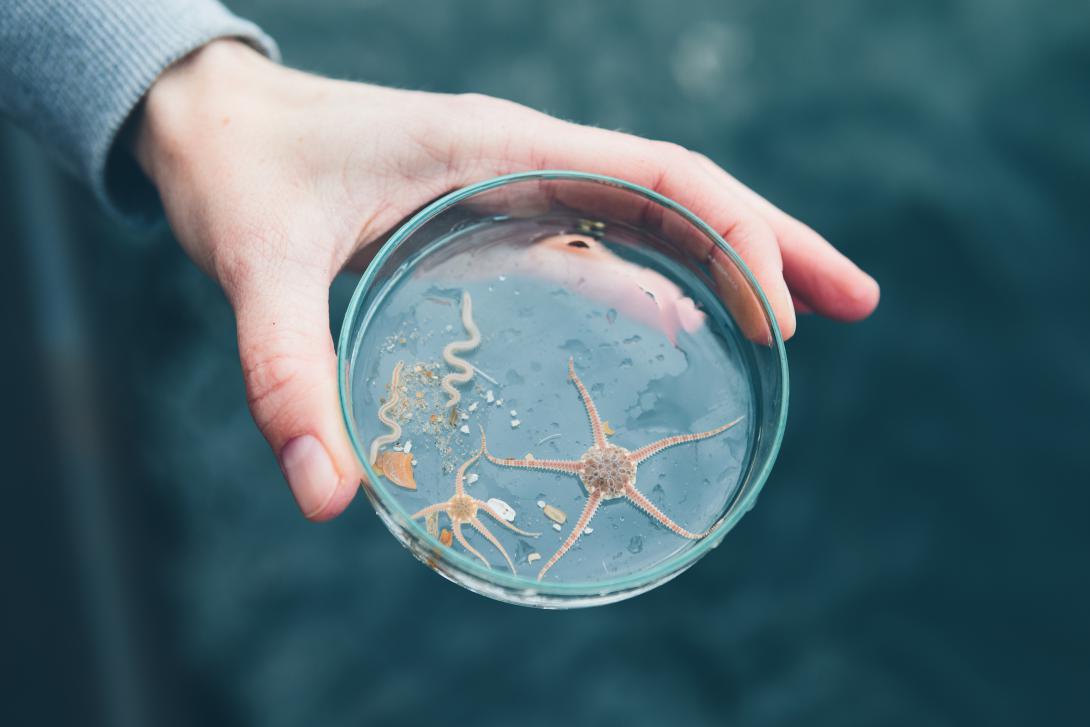

Data & Services
Related stories
Relevant news
-
A Taxon-Match tool for AlgaeBase, based on the WoRMS Taxon Match 
A Taxon-Match tool for AlgaeBase, based on the WoRMS Taxon Match
A taxon-match tool built on AlgaeBase is now available through the LifeWatch eLab, with the same functionalities as the WoRMS taxon-match tool. -
Tracking sharks in the North Sea for better protection and management 
Tracking sharks in the North Sea for better protection and management
With support from LifeWatch Belgium and the European Tracking Network (ETN), researchers are tagging sharks in the Belgian part of the North Sea to uncover their movements and preferred habitats. Scientists from the Flanders Marine Institute (VLIZ) and the Research Institute for Agriculture, Fisheries and Food (ILVO) lead this work, using LifeWatch infrastructure to collect vital data that will help guide targeted protection and management. -
Fish Don’t Know Borders: Tracking Aquatic Life Across Europe 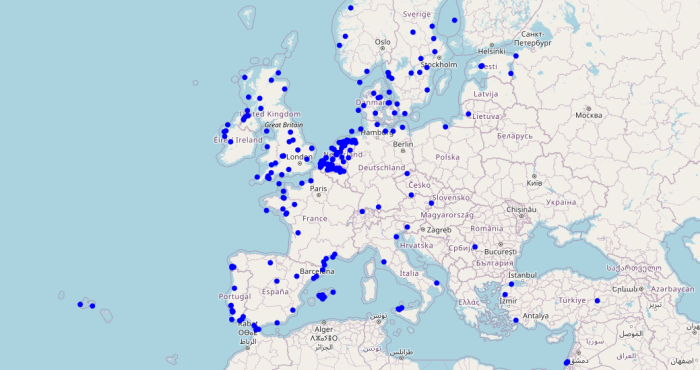
Fish Don’t Know Borders: Tracking Aquatic Life Across Europe
Through the European Tracking Network (ETN), LifeWatch Belgium connects researchers who track fish and other aquatic species across borders. Using a shared network of acoustic receivers, scientists follow animal movements from rivers to seas, revealing how aquatic life links ecosystems throughout Europe. -
From Europe to the Atlantic: new insights into eel migration 
From Europe to the Atlantic: new insights into eel migration
European eels are legendary travelers, undertaking journeys of up to 9,000 km to spawn inthe Atlantic Ocean. A groundbreaking study led by LifeWatch researchers has broughttogether tracking data from more than 2,300 eels across 9 countries, revealing howgeography and river barriers shape the timing of this epic migration.





